DataLad extensions
The core functionality of DataLad software can be broadened by thematic extensions. Extensions are shipped as separate Python packages, and you can browse a list at the DataLad extensions GitHub page.
Development of the following extensions was directly influenced by the SFB:
DataLad-Gooey (GUI)
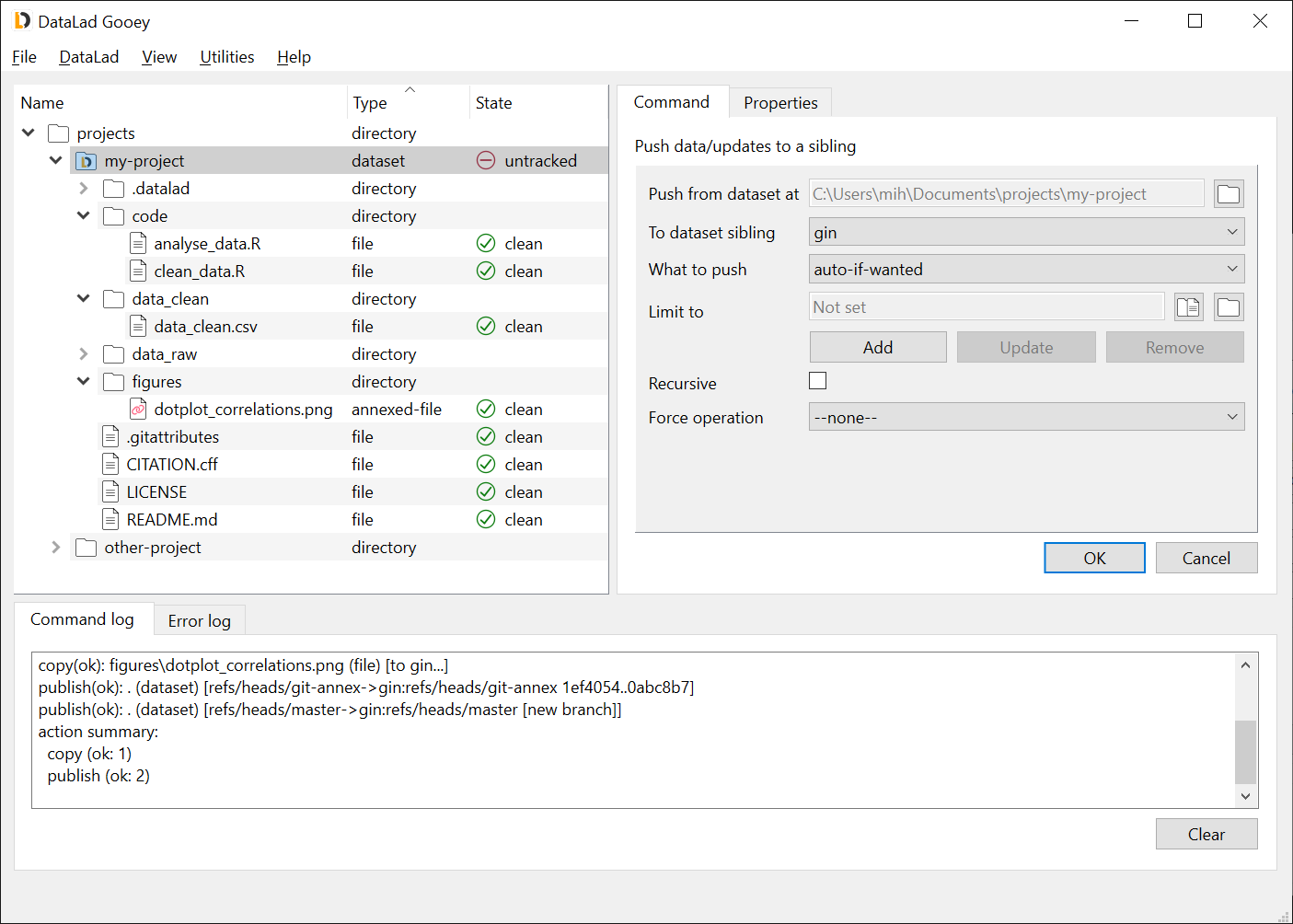
The DataLad-Gooey (pronounce "GUI") package provides a graphical user interface for DataLad. It is specifically aiming at making key data management tasks more accessible and more convenient, without requiring to become familiar with the command line.
While using DataLad Gooey assumes at least some familiarity with DataLad concepts, the simplified command suite makes starting with DataLad easier via tailor-made command selections, condensed parameter specifications, and tool tips.
It comes with a dedicated Windows installer, and can be installed on all major operating systems from PyPI.
Learn more from the DataLad-Gooey documentation.
DataLad-XNAT
XNAT is an open source imaging informatics platform developed by the Neuroinformatics Research Group at Washington University. It facilitates common management, productivity, and quality assurance tasks for imaging and associated data. XNAT can be used to support a wide range of neuro/medical imaging-based projects. Several imaging centres within the SFB1451 are using XNAT.
DataLad can be equipped with datalad-xnat, an extension providing a set of commands to track XNAT projects. The extension features:
- Authenticated and anonymous access to XNAT
- Support for local deployments
- Compatibility with major public deployments: ConnectomeDB, NITRC IR, XNAT Central
- Full or constrained exports of entire instances, individual projects, single subjects, particular acquisitions, and limit by resource collections
- Flexible dataset structuring with support for multiple layers of nesting
- Parallel downloads
You can read more in the DataLad-XNAT github page and in datalad-XNAT documentation.
DataLad-NEXT
DataLad-NEXT is a proving ground for broadly applicable novel functionality.
Most important for SFB is the much improved ability to deposit DataLad datasets on any WebDAV server. This includes UKK's Dracoon system, as well as Sciebo, EUDAT's B2DROP, and self-hosted OwnCloud or NextCloud instances.
The exported content can be made browsable like normal content for non-DataLad consumers, but also remain a full dataset that can be cloned and collaborated on. While it won't be as fast or flexible as with more specialized services for Git or data hosting, this is a great approach for dataset deposition, where only one site is uploading dataset and file content updates, and the data consumers can choose between using DataLad or point-and-click interfaces. See the WebDAV walkthrough page for a detailed overview.
Read more in DataLad-NEXT github page and in DataLad-NEXT documentation.
DataLad-REDCap
DataLad-REDCap extension provides convenience commands for exporting data from REDCap into DataLad datasets. It is yet unreleased, but you can install a working development version from DataLad-REDCap Github and read DataLad-RedCAP documentation.